Volume II ● Issue 1/201 ● Pages 9–14
Bioarchaeological Analysis of Bronze Age Human Remains
from the Podillya Region of Ukraine
Alexey G. Nikitina*
aBiology Department, Grand Valley State University, 1 Campus Drive, Allendale, MI 49401, USA
Abstract
Archaeological discoveries in the Podillya region of Ukraine produced a wealth of pre-historic artifacts, including human remains. When human remains are not accompanied by material culture artifacts their placement in a historical context is difficult, without a comprehensive analysis using a variety of archeometric and bioarchaeological tools. At one archaeological site, connected to the Eneolithic Trypillia-Cucuteni culture complex, two osteological specimens with no material culture context were discovered. Radiocarbon analysis of these remains placed them within the timeframe of a local Bronze Age group. Ancient mitochondrial DNA analysis identified a maternal genetic lineage connection to north-central Europe. Physical anthropology analysis of one of the two specimens revealed evidence of postmortem manipulation. The discovery of Bronze Age remains in an Eneolithic ritual context raises questions about burial practices of regional Bronze-Age inhabitants.
Article info
Article history:
Received: 14 May 2011
Accepted: 13 September 2011
Key words:
Ukraine
Podillya
Verteba Cave
Eneolithic-Bronze Age
Trypillia-Cucuteni culture complex (TC)
mitochondrial DNA (mtDNA)
ancient DNA (aDNA)
1. Introduction
The Podillya region of Ukraine lies 100 km east of the foothills of the Carpathians. Archaeological evidence suggests that human activity in the area began with Acheulean technologies in the lower Paleolithic, and vigorously continued through the remainder of the Pleistocene (Sytnyk 2000; Madeyska 2002). Human occupation of the area continued through the Mesolithic (Mackevyj 1997) and into the Neolithic (Mackevyj, 2001). In the Neolithic the area was home to Linearbandkeramik (LBK) groups of the first European farmers (Konoplya 2008), and, subsequently, to a cultural conglomerate called the Trypillia-Cucuteni complex (TC) (Nikitin et al. 2010).
The crown jewel of Podillyan archaeology is Verteba, a gypsum cave housing remarkable artifacts, including human remains, from the middle to late TC period (Nikitin et al. 2010). When it comes to human remains Verteba presents a puzzling situation. Most human remains in the cave are found in a disarticulated state. Singular bones belonging to one or two individuals along with pottery shards and animal bones, sometimes mixed with charcoal, represent the vast majority of the finds (Nikitin et al. 2010 and unpublished data). These finds could represent secondary burials, or token deposits. The only report of articulated skeletons comes from Stojanowski (1948), where he provides an anthropological analysis of skulls from 21 skeletons found in a corner of the cave where the individuals were apparently killed by a rockslide. However, it is unclear if these are TC remains or they belong to another culture.
Most TC remains along with material culture artifacts found in recent years have been retrieved from the cave’s cultural layers (Nikitin et al. 2010). Occasionally, singular bones or bone assemblages are also found on the cave floor, but their cultural association can be dubious. This article provides a comprehensive analysis of two such bones from Verteba Cave.
2. Materials and Methods
2.1 Specimen collection and examination
Specimens were collected during the 2005 field season. The finds consisted of a human fibula (F3) and vertebra (V6). Both specimens were found by the author on an unexcavated surface along the cave wall, in an area not exposed to human traffic nor archaeological work. The vertebra was physically intact, while the distal end of the fibula was broken off. No material culture artifacts were associated with the specimens. Physical anthropology and forensic pathology analyses of the vertebra were conducted by Dr. Norman J. Sauer at the Forensic Anthropology Lab of Michigan State University. Radiocarbon dating was performed at the Kyiv Radiocarbon Laboratory after all specimen examination, including DNA extraction, was completed. Calibrated dates were obtained using OxCal 4.1.3 online calibration software (Bronk Ramsey 2009), with IntCal 04 atmospheric curve (Reimer et al. 2004).
2.2 Ancient DNA analysis
Precautions were taken from the moment artifacts were sighted to avoid contamination with modern DNA. All sample cleanup, extraction and amplification procedures were conducted by a single person (the author of this report). DNA extraction promptly followed specimen collection, to reduce sample deterioration and any potential exposure to DNA contaminants.
The ancient DNA (aDNA) retrieval and processing adhered to sterile technique standards (Cooper, Poinar 2000) as much as possible. All specimen cleanup and pre-extraction processing took place in a newly-built lab under an exhaust hood used solely for aDNA work. Protective clothing, gloves and facemasks were used at all times. The hood surface was thoroughly wiped with ethanol, bleach and DNA-Erase solution (MP Biomedicals) before and after each use. Gloves were changed with every step of the procedure. All equipment and tools (rotary tool, weigh boats, mortars and pestles) were sterilized prior to use.
Specimen cleanup, DNA extraction and DNA analysis were spatially separated. Prior to DNA extraction, bones were cleaned of surface dirt and exposed to 253.7 nm UV light for 60 min on all sides. Specimens were subsequently cleaned by sanding off ~1 mm of the surface with a Dremel rotary tool. Cleaned bones were exposed to UV light for an additional 20 min after sanding. Thereafter, 300–400 mg of bone were cut from each specimen with a cutting disk and ground into fine powder with a porcelain mortar and pestle.
DNA was extracted from each specimen in two temporally separated extractions, in separate laboratory facilities dedicated to aDNA work, under a laminar flow hood with a built-in UV source. Powdered bone material was washed three times with sterile EDTA, pH 8.0, and rinsed with sterile molecular grade water, pH 7.5–8.0. DNA was then extracted following a modified Qiagen protocol for extraction of DNA from bone (Qiagen Inc., Valencia, CA, USA). DNA was eluted with 50 µl of sterile water, pH 8.5, and frozen at –20 oC. DNA extraction from each specimen was accompanied by blank extractions and separate controls for each reagent used in the extraction.
Ancient DNA was amplified using the Polymerase Chain Reaction (PCR) procedure. The PCR setup took place under a laminar flow hood solely dedicated to aDNA PCR work. The thermocycler used to perform all PCRs was spatially separated from the rooms where the bone preparation and DNA extraction procedures took place. All amplifications were done using a Qiagen FastCycle PCR kit following the conditions outlined in the kit protocol. PCR reactions were prepared with 0.2–2 µl of the template. The initial PCR amplification for each specimen extract contained a positive control to detect residual PCR inhibitors.
The haplogroup status of each specimen was determined by amplifying and analyzing diagnostic segments of the mtDNA coding region and the first hypervariable segment (HV1) of the control region of mtDNA. Amplifications were carried out until four independent PCR products for each amplified fragment per extraction for each sample were obtained.
Coding region mtDNA fragments containing diagnostic polymorphisms for the HV clade (a T-C transition at nucleotide 14766 producing a non-polymorphic Mse I restriction pattern) and the H haplogroup (a 7028T-C transition producing a non-polymorphic Alu I restriction pattern) were amplified for both specimens. In addition, a diagnostic region for the U/K clade around an A-G transition at nucleotide 12308 producing a polymorphic Hinf I restriction pattern was amplified for the fibula extracts. Coding region amplifications were performed using previously published primers (Santos et al. 2004).
The HV1 region was amplified in two overlapping segments, 235 bp and 245 bp, produced by the following primers:
L15993 (ACTCCACCATTAGCACCCAA),
H16189 (CTTGCTTGTAAGCATGGGGA),
L16185 (AACCCAATCCACATCAAAACC),
and H16391 (AGGATGGTGGTCAAGGGAC). The second fragment incorporated the author’s HV1 polymorphism at nucleotide 16304, thus adding a one-step contamination control in the phylogenetically informative 16209–16356 “middle region” of HV1 (Gilbert et al. 2003). HVI primers were designed using the NCBI/Primer-BLAST Primer Designing Tool (http://www.ncbi.nlm.nih.gov/tools/primer-blast/index.cgi).
Amplification products of the coding region were subjected to Restriction Fragment Length Polymorphism (RFLP) analysis. Nucleotide positions producing specific RFLP patterns were confirmed by direct sequencing of the amplicons. The HV1 amplification fragments were directly sequenced. The DNA sequence analysis was done at the University of Michigan DNA Sequencing Core. All sequencing chromatograms were visually inspected. Sequence analysis was accomplished using NCBI BLAST (http://blast.ncbi.nlm.nih.gov/Blast.cgi).
3. Results
3.1 Carbon dating, physical anthropology and forensic pathology analyses
Radiocarbon dating assigned both bone specimens to approximately 8th century BCE (Table 1). The dates place the remains within the timeframe of the Bronze-Age Gáva-Holigrady culture, an eastern Hallstatt group (Smirnova 1976). The culture existed from 10–7th century BCE and is thought to have had Thracian influences (Grakov et al. 1977). Holigrady, the eponymic center of the culture, is just 15km south from Verteba.
The vertebra possesses two demifacets on both sides of the body indicating that it is thoracic, and identified as between T2 and T8. The superior and inferior surfaces of the body showed billowing and no evidence of epiphyseal ring fusion. There were deep groves on the superior and inferior surfaces at the attachment sites between the neural arch and the body. These remnants of deep grooves indicate evidence of recent fusion between the neural arch and the vertebral body.
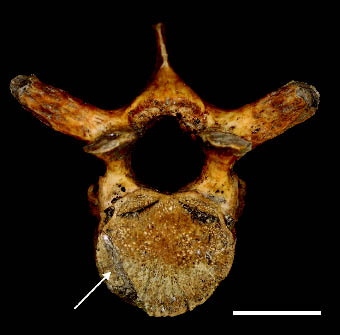
On the superior surface of the vertebral body there was another deep groove consistent with sharp force trauma crossing anterior-medially to posterior-laterally (Figure 1). This groove showed no discoloration, suggesting that the damage was post-mortem but not related to recovery.
The V6 specimen was compared to Dr. Sauer’s laboratory collection of specimens of known age. This comparison suggested that the vertebra from Verteba Cave is from an individual who died between 10 and 16 years of age.
The physical anthropology analysis of the fibula determined that it belonged to a full adult. Thus, the two osteological samples came from two different individuals.
3.2 Ancient DNA analysis
Coding region nucleotide analysis of the vertebra produced a polymorphism pattern characteristic for mtDNA haplogroup H. The HV1 sequence analysis for the specimen revealed an unambiguous transition at nucleotide 16070, relative to the mtDNA Cambridge Reference Sequence (rCRS; Anderson et al. 1981; Andrews et al. 1999) (Table 1).
The HV1 region analysis of the fibula revealed transitions at nucleotide positions 16256 and 16270, placing this specimen in the U5a1a subhaplogroup of the U clade (Richards et al. 1998) (Table 1). Since position 16270 is considered to be highly susceptible to postmortem modification in aDNA samples (Gilbert et al. 2003), caution was taken when dealing with this position. All four 245-bp HV1 amlicons from the second extraction and three out of four amplicons from the first extraction of the F3 specimen showed an unambiguous 16256–16270 motif. Thus the 16256–16270 motif was considered endogenous to the F3 specimen. The coding region RFLP analysis of mtDNA of the fibula was consistent with the placement of the specimen in the mtDNA haplogroup U (Table 1). DNA sequences have been deposited in GenBank (http://www.ncbi.nlm.nih.gov/genbank/) under accession numbers JN636826 (F3) and JN636827 (V6).
4. Discussion
Previous studies determined that human ritual activity at Verteba Cave was restricted to TC groups and ceased after the Eneolithic (Nikitin et al. 2010). This report extends the possible use of the cave into the Bronze Age. While the two osteological specimens described in this report are not associated with a cultural context, the radiocarbon analysis places them within a timeframe for a specific local Bronze Age group while the aDNA analysis puts them in the local European context as well.
Both Bronze Age bone specimens described here were found on the cave floor. It is difficult to determine how such surface finds connect to the human exploitation of the cave. Perhaps such artifacts ended up in the cave by accident, but it is also possible they could have been intentionally deposited during ancient times. The location of the trauma on the superior surface of the vertebra can be viewed as evidence of a secondary burial practice (N. Sauer, personal communication, 2005). At the same time, it can also be a tooth mark left by an animal (G. Madden, personal communication, 2008). In the latter case, the bone may have been dragged into the cave from the surface, or carried with alluvial deposits. At this time, it is impossible to make a distinction among these possibilities.
The vertebra’s mtDNA haplogroup H is the most common haplogroup in Europe. The 16070 polymorphism found in the vertebra has been identified in H-bearing Hutterites (Pichler et al. 2010) who trace their origins from the Tyrol province in Austria, as well as in Finns (Finnilä et al. 2001) and Slovenians (Zupanič Pajnič et al. 2004). Beyond these associations it is difficult to track the distribution of this particular mtDNA variant. However, it is worth noting that mtDNA lineages of the H haplogroup were uncovered in the Eneolithic remains at Verteba as well (Nikitin et al. 2010), suggesting regional maternal genetic lineage commonality.
The Bronze Age fibula’s mtDNA lineage belongs to the U5a1a branch of the U5 subclade of the U clade characterized by a back mutation at position 16192 (Richards et al. 1998). U5 is considered a European-specific mtDNA subclade of an estimated age of ~50,000 years, making it the oldest European mitochondrial lineage that can be traced back to the first modern human settlers of Europe (Richards et al. 1998). The U5a1a sublineage is also found in the Near East and, thus, could have been migrating in and out of Europe (Richards et al. 2000). This sublineage is common in pre-historic European remains. Specimens harboring the 16256–16270 HV1 motif have been found in the Neolithic LBK/AVK remains (Haak et al. 2010) as well as in the late Chalcolithic remains from northeast Poland (Burger et al. 2007). When the ancient and modern occurrences of U5a1a in west Eurasia are plotted on a regional map, they appear to cluster in north-central Europe, diffusing towards the Ponto-Caspian region, placing the Verteba Cave specimen in the center of that gradient (Figure 2). Thus, maternal genetic lineages of both Bronze Age specimens from Verteba seem to have local central European roots.
The current archaeological opinion is that the Bronze Age cultures of Ukraine are not connected to Eneolithic TC groups (M. Videiko, personal communication, 2009). To date, Bronze Age material culture remains at Verteba Cave are represented by only a few bronze items including a spear, an axe, and several beads (Ossowski 1895). The presence of human remains from the Bronze Age within the overall Eneolithic context of the cave is intriguing. Unfortunately we cannot determine if the Bronze Age bones were purposefully deposited in the cave. Since the age difference between the cave’s oldest human remains from TC (Nikitin et al. 2010) and the Bronze Age remains is about 2,200 years, any ritual continuity between Eneolithic and Bronze Age populations around Verteba would be remarkable indeed. As only about 25% of the cave has been explored to date, further examination of the cultural and biological artifacts at Verteba should help clarify the extent of the cave’s ritual use by the cultures that succeeded Trypillia.
Acknowledgements
The author would like to thank Mykhailo Sokhatsky, Director of Borschiv Regional Museum and Curator of Verteba Cave, for access to the cave’s artifacts, as well as Ian McBain, Catherine Willis and Cynthia June for their assistance with DNA analysis. Special thanks to Dr. Norm Sauer for the physical anthropology analysis of the osteological remains and to Jessica Riley for critical reading and thoughtful comments on the manuscript. This work was supported, in part, by a Research Seed Grant from the Michigan Space Grant Consortium and by research funds from GVSU.
References
Anderson, S., Bankier, A. T., Barrel, B. G., de Bruijn, M. H. L., Coulson, A. R., Drouin, J., Eperon, I. C., Nierlich, D. P., Roe, B. A., Sanger, F., Schreier, P. H., Smith, A. J. H., Staden, R., Young, I. G. 1981: Sequence and organization of the human mitochondrial genome. Nature 290, 457–465.
Andrews, R. M., Kubacka, I., Chinnery, P. F., Lightowlers, R. N., Turnbull, D. M., Howell, N. 1999: Reanalysis and revision of the Cambridge reference sequence for human mitochondrial DNA. Nature Genetics 23, 147.
Bronk Ramsey, C. 2009: Bayesian analysis of radiocarbon dates. Radiocarbon 51, 337–360.
Burger, J., Kirchner, M., Bramanti, B., Haak, W., Thomas, M. G. 2007: Absence of the lactase-persistence-associated allele in early Neolithic Europeans. Proceedings of the National Academy of Sciences USA 104, 3736–3741.
Cooper A., Poinar, H. 2000: Ancient DNA: Do it right or not at all. Science 289, 1139.
Costa, M. D., Cherni, L., Fernandes, V., Freitas, F., Ammar el Gaaied, A. B., Pereira, L. 2009: Data from complete mtDNA sequencing of Tunisian centenarians: testing haplogroup association and the “golden mean” to longevity. Mechanisms of Ageing and Development 130, 222–226.
Finnilä, S., Lehtonen, M. S., Majamaa, K. 2001: Phylogenetic network for European mtDNA. American Journal of Human Genetics 68, 1475–1484.
Gilbert, M. T. P., Willerslev, E., Hansen, A. J., Barnes, I., Rudbeck, L., Lynnerup, N., COOPER, A. 2003: Distribution patterns of postmortem damage in human mitochondrial DNA. American Journal of Human Genetics 72, 32–34.
Grakov, B. N., Yelagina, N. G., Yatsenko, I. V. 1977: Early Iron Age. Moscow University Press, Moscow.
Haak W., Balanovsky O., Sanchez J. J., Koshel S., Zaporozhchenko V., et al. 2010: Ancient DNA from European Early Neolithic Farmers Reveals Their Near Eastern Affinities. PLoS Biology 8(11), e1000536.
Konoplya ,V. 2008. The settlement of Linear Pottery Culture Blyschanka at the Seret River. Proceedings Institute of Archaeology of L’viv University 3, 196–222.
Madeyska, T. 2002: Evidence of climatic variations in loess and cave Palaeolithic sites of southern Poland and western Ukraine. Quaternary International 91, 65–73.
Mackevyj, L. G. 1997: On the Ukrainian Carpathian Mesolithic. Third Annual Meeting of the European Association of Archaeologists. Ravenna, 1997, p. 38.
Mackevyj, L. G. 2001: Das Mesolithikum der äußersten Westukraine. From the Mesolithic to the Neolithic. Budapest, 2001, pp. 313–337.
Malyarchuk, B., Derenko, M., Grzybowski, T., Perkova, M., Rogalla, U., et al. 2010: The Peopling of Europe from the Mitochondrial Haplogroup U5 Perspective. PLoS ONE 5(4), e10285.
Nikitin, A. G., Sokhatsky, M. P., Kovaliukh, M. M., Videiko, M. Y. 2010: Comprehensive site chronology and ancient mitochondrial DNA analysis from Verteba Cave -a Trypillian culture site of Eneolithic Ukraine. Interdisciplinaria Archaeologica 1, 9–18.
Pichler, I., Fuchsberger, C., Platzer, C., Çalişkan, M., Marroni, F., Pramstaller, P. P., Ober, C. 2010: Drawing the history of the Hutterite population on a genetic landscape: inference from Y-chromosome and mtDNA genotypes. European Journal of Human Genetics 18, 463–470.
Ossowski, G. 1895: Sprawozdanie czwarte z wycieczki paleoetnologicznej po Galicyi w roku 1892. Zbior Wiadomosci do Antropologii Krajowej 18, 1–28.
Reimer, P. J., Baillie, M. G. L., Bard, E., Bayliss, A., Beck, J. W., Bertrand, C. J. H. et al. 2004: IntCal04 terrestrial radiocarbon age calibration, 0–26 cal kyr BP. Radiocarbon 46, 1029–1058.
RICHARDS, M. B., MACAULAY, V. A., BANDELT, H.-J., SYKES, B. C. 1998: Phylogeography of mitochondrial DNA in western Europe. Annals of Human Genetics 62, 241–260.
Richards, M., Macaulay, V., Hickey, E., Vega, E., Sykes, B., Guida, V., Rengo, C., Sellitto, D., Cruciani, F., Kivisild, T., et al. 2000: Tracing European founder lineages in the Near Eastern mtDNA pool. American Journal of Human Genetics 67, 1251–1276.
Santos, C., Montiel, R., Anglés, N., Lima, M., Francalacci, P., Malgosa, A., Abade, A., Aluja, M. P. 2004: Determination of human Caucasian mitochondrial DNA haplogroups by means of a hierarchical approach. Human Biology 76, 431–453.
Schoenberg, A., Theunert, C., Li, M., Stoneking, M., Nasidze, I. 2011: High-throughput sequencing of complete human mtDNA genomes from the Caucasus and West Asia: high diversity and demographic inferences. European Journal of Human Genetics 19, 988–994.
Smirnova, G. I. 1976: The remains of Gava-Goligrady in the eastern basin of the Carpathians. Archeologicheskij Sbornik 17,18–34.
Stojanowski, K. 1948: Antropologia prehistoryczna Polski. Polska Akademia Umiejetnosci, Kraków.
Sytnyk, O., 2000: Middle Palaeolithic of Podilia. National Ukrainian Academy of Sciences, Lviv.
Zupanič Pajnič, I., Balažic, J., Komel, R. 2004: Sequence polymorphism of the mitochondrial DNA control region in the Slovenian population. International Journal of Legal Medicine 118, 1–4.
*Corresponding author, e-mail: nikitin@gvsu.edu
Table 1. Radiocarbon dates and mtDNA sequence analysis, including the status of the coding region diagnostic RFLPs and control region (HV1) nucleotide polymorphisms, for Bronze Age human remains from Verteba Cave and the mtDNA polymorphism status of the author of the report. The HV1 variation denotes nucleotide transitions relative to the Cambridge Reference Sequence (rCRS) (Anderson et al. 1981; Andrews et al. 1999).
Specimen Uncalibrated date, Calibrated date, Coding region RFLPs Control region Haplogroup
BP (laboratory code) cal BC (95.4%, 2 ∂) (HV1) variation
Fibula (F3) 2,610±80 (Ki 13387) 969–418 14766 Mse I+ 16256, 16270 U5a1a
12308 Hinf I+
7025 Alu I+
Thoracic vertebra (V6) 2,660±80 (Ki 13386) 1,013–544 14766 Mse I– 16070 H
7025 Alu I–
Author 14766 Mse I– 16304 H5a
12308 Hinf I–
7025 Alu I–
Figure 1. Superior view of thoracic vertebra from Verteba Cave with arrow pointing to surface trauma. Image courtesy of Norman J. Sauer, Michigan State University, 2005.
Figure 2. Map of west Eurasia showing the distribution of the U5a1a mtDNA haplotype. Black circles indicate ancient specimens (data from Burger et al. 2007; Haak et al. 2010; this report); red circles represent modern samples (data from Costa et al. 2009; Malyarchuk et al. 2010; Schoenberg et al. 2011). The location of the U5a1a specimen from this report is outlined with a square.
0 3 cm
